Warning! The missing data problem
The VSNi Team
12 May 2021
Missing data are a common problem, even in well-designed and managed research studies. It results in a loss of precision and statistical power, has the potential to cause substantial bias and often complicates the statistical analysis. As data analysts, it is crucial that we understand the reasons for the missing data and apply appropriate methods to account for it. Failure to do so leads to trouble! Let’s consider an example.
The following figure illustrates data from a study on the intelligence quotient (IQ) of students living in a city, town, or village. A random selection of students was recruited into the study and their IQs measured. Unfortunately, IQ measurements were not available for all students in the study, resulting in missing data (denoted by NA). To make valid inferences, it is very important that we understand why the data are missing, and what we can do about it.
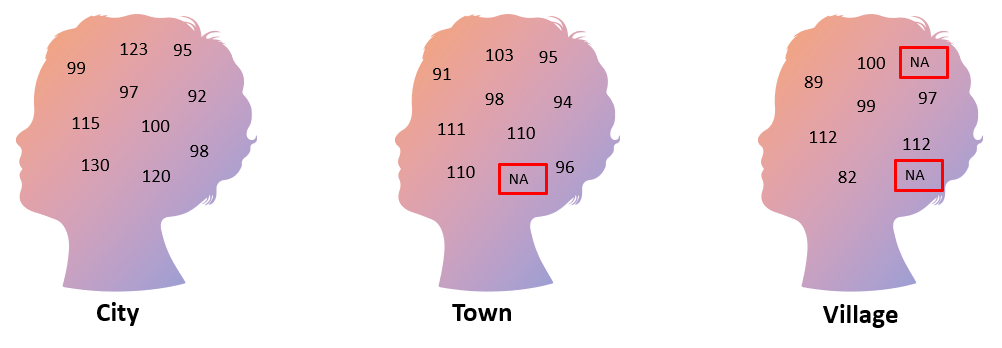
There are three main types of missing data:
Missing completely at random (MCAR)
Here missingness has nothing to do with the subject being studied: instead the missing values are an entirely random subset of the data. In our example, the missingness could be considered MCAR if the missing IQ test results were accidentally deleted by the researcher.
Missing at random (MAR)
Here missingness is not related to the value of the missing data but is related to other observed data. For example, we might find that younger students were less likely to complete the IQ test due to some factor unrelated to their IQ, such as a shorter attention span than older students. This missing data can be considered MAR, as failure to complete the test had nothing to do with their IQ (after accounting for age).
Missing not at random (MNAR)
When the missing data are neither of the above, they are MNAR. In other words, missingness is related to the value of the missing data. For example, if failure to complete the IQ test was related to the student’s IQ the missingness would be MNAR.
When the data are MCAR, missingness doesn’t induce bias. However, this isn’t the case for MAR and MNAR, and great care needs to be taken to ensure that this does not lead to biased and misleading inferences.

Let’s go back to our example. Assuming the MCAR mechanism, we could analyse the data in Genstat using an unbalanced ANOVA. Assuming that MAR is conditional on age, an unbalanced ANOVA with age as a covariate would be an appropriate analysis. But what to do if the missingness is MNAR? This is much more problematical. Indeed, the only way to reduce potential bias is if we can explicitly model the process that generated the missing data. Challenging indeed!
As our example illustrates, when faced with missing data we must apply an appropriate statistical method to accommodate it. The choice of method will depend on the nature of the missingness, in addition to the type of data we have, the aims of our analysis, etc.
But what about imputation?
Imputation replaces missing values with estimated values. That is, the dataset is completed by filling in the missing values. Many different methods of imputation exist, including mean substitution, regression imputation, EM algorithm, and last observation carried forward. Be warned however, imputation may not overcome bias - indeed it may also introduce it! In addition, it does not account for the uncertainty about the imputed missing values, resulting in standard errors that are too low. However, if the proportion of missing values in the dataset is small, imputation can be useful. Many statistical methods can handle datasets with missing values e.g., maximum likelihood, expected maximization, Bayesian models. Others, such as principal component analysis and some spatial or temporal mixed models, require complete datasets. Imputation allows us to apply statistical methods requiring complete datasets.
As a final thought - the best possible solution for missing data is to prevent the problem from occurring. Carefully planning and managing your study will help minimize the amount of missing data (or at least ensure it is MCAR or MAR!). When you have missing data, use an appropriate statistical technique to accommodate it. Statistical or computational techniques for imputing missing data should be the last resort.
Popular
Related Reads
