Accelerating sugar beet breeding insights for SESVanderHave
Accelerating sugar beet breeding insights for SESVanderHave
Analysing large phenotypic and genotypic datasets using the H3-pipeline
Belgium-based SESVanderHave stands tall as a global leader in sugar beet breeding, selling billions of seeds annually. With their commitment to innovation and quality, they have successfully developed localised sugar beet varieties for individual markets in over 50 countries.
The company uses a variety of cutting-edge technologies, including the H3-pipeline, to develop new varieties of sugar beet that have higher yields and improved resistance to pests and diseases.
Managing the challenge of big data in plant breeding
SESVanderHave faced significant challenges due to the exponential growth of data in plant breeding. With new candidate varieties being tested each season, characterised in greater detail and incorporating phenotypic and molecular marker data, they needed a powerful solution that could model and condense this vast amount of information for effective decision making.
A new statistical tool for sugar beet breeding
Recognising these challenges, the H3 consortium, comprising VSNi, Biometris and JHI, collaborated with SESVanderHave to develop a cutting-edge software pipeline. This innovative pipeline seamlessly integrates data and models, enabling breeders to obtain relevant information essential for their work. The pipeline encompasses various features, including the design and analysis of phenotyping experiments, allele transmission from parents to offspring, coupling of phenotypes to genotypes in QTL mapping and GWAS models, and genomic prediction models.
One of the key requirements from SESvanderHave was the ability to analyse vast phenotypic and genotypic datasets swiftly. The H3-pipeline needed to exhibit exceptional speed and robustness: the DNA of thousands of young plants are genotyped, and based on advanced genomic prediction methods the best new genotypes are selected and field tested in numerous global locations to ensure their performance under diverse conditions.
The H3-pipeline: essential components and processes
The H3-pipeline is a crucial component of the breeding program at SESVanderHave, comprising different modules that cover key steps in the selection of lines. The process begins with examining the trial field layout and generating an optimal experimental design using R. Next, the pipeline analyses field trials using the SpATS R package to calculate Best Linear Unbiased Estimates (BLUEs) and heritabilities and identify potential outliers. Subsequently it creates detailed reports for each trial, summarising the results for all genotypes and their important traits. Next it computes Best Linear Unbiased Predictions (BLUPs) across environments using ASReml-R, while employing genomic selection to identify new genotypes based on a comprehensive training set. Additionally, the pipeline incorporates a crossing tool that uses genomic predictions to identify the ideal parent combinations for the upcoming breeding cycle.
The breeding pipeline's analytics were developed by Biometris in the R programming language using the ASReml library. To ensure flexibility and modularity, VSNi implemented the pipeline using a microservices architecture. This approach allows independent development and deployment of different modules within the pipeline.
For efficient management and execution of the pipeline's modules, JHI developed a dashboard using a combination of Java and Vue.js. The dashboard is designed to run seamlessly in Docker environments, allowing it to be easily scaled up as required.
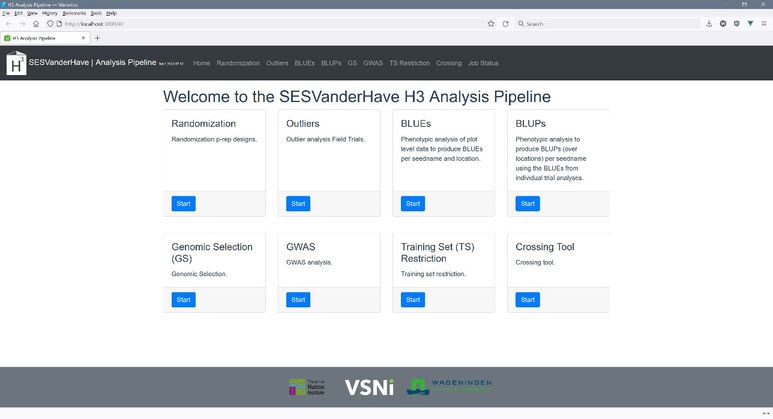
The dashboard of the SESVanderHave H3 Analysis Pipeline
Higher yields, improved disease resistance
Agriculture is currently undergoing rapid and profound changes, such as the consumer-driven reduction in demand for plant protection products and fertilisers. This trend is reinforced by climate change, which has already led to an expansion of the occurrence of plant pathogens. In addition, climate change causes more extreme weather conditions, such as extended heat and drought events with significant impacts on crop yields. This plethora of changes to the agricultural context confronts breeding companies like SESVanderHave with an abrupt market-demand shift towards novel crop varieties with a higher level of tolerance and/or resistance against a wide range of pathogens as well as environmental stresses like drought and heat. At the same time, yield and agricultural output must remain at a competitive level to maintain profitability across the value chain.
SESVanderHave’s vision for the development of modern sugar beet varieties is to enable the prediction of variety performance in both known and unknown environmental conditions. This predictive ability relies on a combination of genotypic data, environmental data and a priori physiological knowledge derived from yield trait components, most of which are obtained using new image capturing technologies.
Over the past ten years, SESVanderHave has intensified its focus on data-driven and predictive breeding, integrating different types of data in the selection process. This has been made possible due to the reduced cost per molecular data point, advancements in phenomics, and the availability of real-time weather data. As a result, we have been able to implement a predictive breeding strategy that allows us to estimate the future performance of our varieties in different target environments.
Hendrik Tschoep, Director of Breeding at SESVanderHave explains their direction:
“It was clear to us that the main challenge was not generating new data but being able to extract the meaningful information needed to maximise the genetic gain in our breeding programs. Therefore, when we began our collaboration with the H3 consortium one of the main challenges was to develop a robust, user friendly, semi-automated statistical pipeline that would allow our breeders to integrate the most relevant data to select the optimal varieties of the future. The H3 pipeline makes it extremely simple, allowing us to utilise different types of data across years, locations, and experiments to select the truly superior offspring in our breeding nurseries.”
The H3 pipeline has helped SESVanderHave to develop new varieties of sugar beet that have higher yields and improved resistance to disease and pests. As a result, SESVanderHave has been able to accelerate the genetic gain for sugar yield coupled with the introgression of new disease traits at the same time, responding to market demands and improving profitability.
Shaping the future
SESVanderHave's collaboration with H3 Consortium has paved the way for groundbreaking advancements in sugar beet breeding. The implementation of the H3 pipeline has empowered SESVanderHave to overcome challenges, develop superior varieties, and shape the future of sugar beet breeding.
“At SESVanderHave we are committed to providing high-quality, sustainable and profitable varieties to our customers. The H3 pipeline does the heavy analytical lifting for us so we can focus on the correct interpretation of the data and selection of the varieties of the future.”
Hendrik Tschoep, Director of Breeding, SESVanderhave
Juan Vegas, senior breeder at SESVanderHave added,
“I’m a sugar beet breeder with more than 10 years of experience at SESVanderHave, and thanks to the new statistical H3 pipeline I’m able to combine different types of data easily, run my own predictions and even simulate new crosses without having to ask our data scientists. It just gives me a lot of freedom and new insights within my breeding programs, helping to unravel the true potential of our genetics.
After years of close collaboration with the H3 consortium, we were able to solve the scientific challenges in breeding data analysis thanks to the long-standing experience of our partners in this field. Most importantly, H3 was able to translate the scientific findings into an analysis pipeline that is stable, reliable and requires minimum intervention, serving a commercial breeding program which has different requirements than tools used in academic science.”

| Would you like to explore how our consultancy services can drive results for your breeding programme? For more information or to dive deeper into the power of H3 Analytics, visit https://vsni.co.uk/services/h3-analytics. |
