Overcoming the Complexity of Modelling Spatial Variation
The VSNi Team
27 March 2024
Genstat's Automatic REML Tool Simplifies Field Trial Analysis
Agricultural variety trials are expensive undertakings, costing significant time and resources. Maximising the return on this investment hinges on rigorous and accurate analysis of field trial data. However, the sheer complexity and scale of these experiments pose considerable challenges. Spatial variation, in particular, adds layers of intricacy, requiring advanced statistical methods to accurately model.
The challenge of spatial variation
In large field trials, spatial variation can significantly impact the accuracy of varietal comparisons. This variation can manifest in various forms, such as soil fertility gradients, micro-climatic effects, and uneven pest or disease pressure. Ignoring these spatial variations can lead to biased estimates, reduced statistical power, and ultimately, suboptimal decision-making.
Restricted Maximum Likelihood (REML) analyses offer a robust solution by enabling us to model the random variation caused by spatial effects. However, deciding upon the best spatial model for each trial is a significant challenge in itself. This complexity lies in the multitude of potential spatial models and correlation structures that can be employed. Factors such as the experimental design, the pattern of spatial variation, and the underlying assumptions must be carefully considered. Navigating this plethora of possibilities can be time-consuming and technically demanding, even with user-friendly menu systems like those offered by Genstat.
Moreover, the optimal spatial model may vary across trials, sites, and experimental conditions. Researchers must navigate this intricate landscape, evaluating various models and selecting the one that best captures the underlying spatial patterns for each individual trial. This process can be repetitive and tedious, particularly when dealing with a large number of trials or experimental sites.
Automating the modelling of spatial variation
A standout feature of Genstat is its Automatic REML Analysis, which tackles the complexity of REML head-on by streamlining the process of modelling spatial variation. Through this innovative approach, the burden of manually exploring and evaluating numerous spatial models is alleviated. Genstat guides users through the analysis process with automated procedures and user-friendly menus, helping them select the best REML model for each trial.
This powerful tool leverages advanced algorithms to identify the optimal spatial model for each trial, ensuring accurate and reliable results, free from the biases and uncertainties that can arise from ignoring or improperly accounting for spatial variation. By automating the intricate model selection process, Automatic REML Analysis enables researchers to model spatial variation effectively and efficiently, without the need for extensive statistical expertise or time-consuming manual efforts.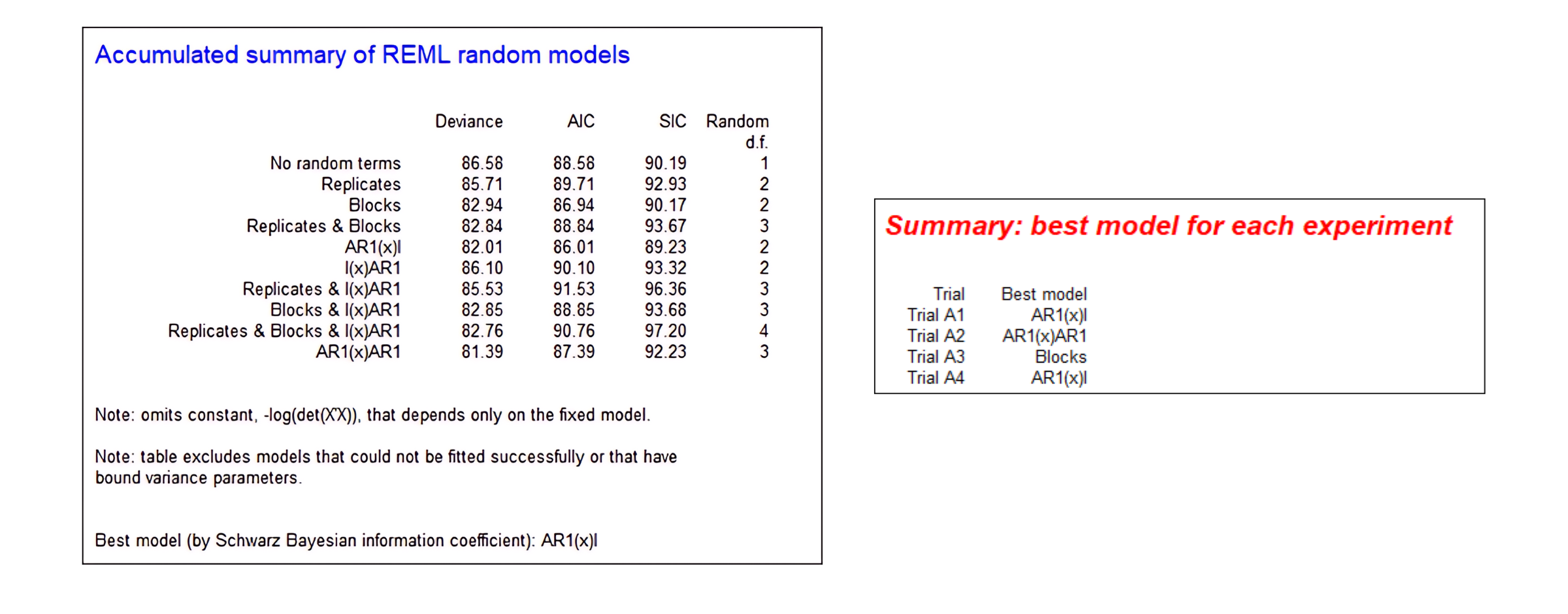
Real-world impact
The practical benefits of Genstat's Automatic Spatial Analysis are profound. Results from 10 years of cereal variety trials in New Zealand with 8-22 varieties per trial and 4 replicates per variety, are insightful. On average, integrating Genstat’s advanced REML methods has given an impressive 20% reduction in Least Significant Differences (LSDs) compared to traditional randomized block analyses.
This substantial reduction in LSDs translates to a significant enhancement in the ability to detect real differences between varieties, offering greater precision and insight into their relative performance. By accurately accounting for spatial variation and capturing the underlying patterns within the trial data, Genstat enables researchers to make more informed decisions and better evaluate the true potential of new and existing crop varieties.
Benefits of Genstat’s Automatic Spatial Analysis
Improved Accuracy: By modelling the changing correlations between plots based on distance, Automatic Spatial Analysis provide more accurate estimates of varietal differences compared to traditional randomized block analyses.
Streamlined Workflow: Genstat's menus guide you through the analysis process for individual trials, whether laid out as block or row-and-column designs, eliminating the need to manually explore all possible model configurations.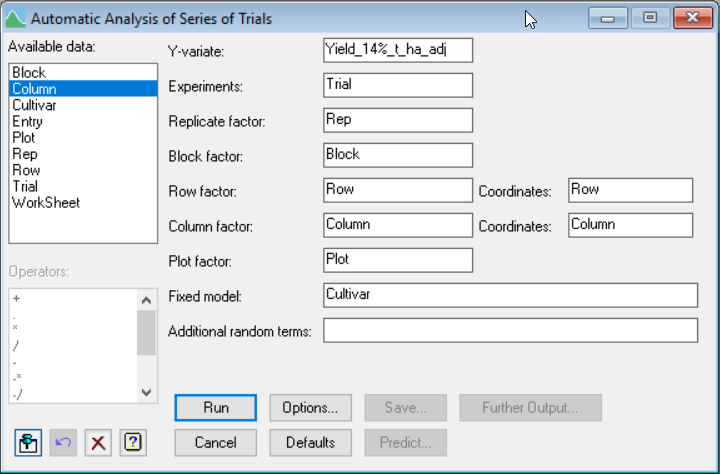
Comprehensive Meta-Analysis: Once you have completed a series of trials across multiple sites, Genstat's meta-analysis menu determines the best models for each trial and combines them into a unified analysis, providing overall estimates of varietal effects and genotype-by-site interactions.
Stability and Interaction Insights: Additional menus allow you to investigate genotype-by-site interactions, assess the site-to-site stability of genotypes, and visualize interactions through GGE biplots.
Seamless Integration for End-to-End Analysis: Genstat's Automatic REML Analysis is more than just a standalone feature; it seamlessly integrates with a powerful suite of tools designed to support researchers throughout the entire data analysis and decision-making process.
You can watch more about Genstat’s Automatic spatial analysis using REML here: https://www.youtube.com/watch?v=gUehFNNvwH0&list=PLfr9hwX9bzpAp92Co8l4XvUrma3Igqb-l&index=13&ab_channel=VSNInternational
For plant breeders and crop scientists, Genstat's Automatic Spatial Analysis provides a powerful way to consistently derive maximum insight from spatial field trial data without specialised expertise. By automating optimal spatial modelling, it unlocks the full potential of these experiments to drive successful variety development and strategic decision-making. With Genstat, breeders can navigate the complex landscape of field trials with confidence, knowing that they have a powerful ally in their quest for innovation.
Make Data-Driven Decisions with Confidence
Get the insights you need to make informed variety selection choices. Explore Genstat or request a quote today!
Popular
Related Reads

